An Overview of Clinical Genetics and its Future
- Rose Begg

- Sep 10, 2020
- 9 min read
Updated: Jul 16, 2023
What is genetic testing?
Geneticists have always been interested in the interplay between genes and characteristic traits.
Genetic diseases are caused by a change or changes, called mutations, in a person’s genome. Most genetic variation doesn’t impact a person’s health. Though sometimes, it only takes one base pair change, out of the approximate 3 billion base pairs which make up our whole genome, to cause a disease.
So in essence, clinical genetics is the study of genetics with the aim to improve disease diagnosis and treatment.
When is genetic testing used?
Currently, the main types of genetic tests check for abnormalities in genes or specific sections of DNA, in chromosome size or number, and in the level and activity of proteins (which itself could also be a sign of genetic abnormality!). Genetic diagnostic testing began in paediatrics, which deals with children’s diseases, but has since gone on to also help diagnose conditions that only arise in adults.
The choice of timing and type of test can be very important. For example, newborn babies can be screened just days after birth for genetic diseases. Prenatal (i.e. before birth) testing can be carried out on a fetus before they are born. However, if potential parents have a specific disease run in their family, they may be carriers. Carriers are healthy people with no disease symptoms but can still pass on a disease-causing mutation onto their children. Potential parents can have genetic testing for this before they even conceive a child.
If a disease is caused by a recessive mutation, such as cystic fibrosis, both genome copies (each from one parent) must contain the mutation. A carrier possesses only one mutated copy and one normal copy of a gene, and so could pass on the mutated gene whilst themselves being completely healthy. If both parents are carriers there is a serious risk of their children inheriting two mutated copies of a gene and so having the disease (see figure 1). Therefore, carrier testing can give future parents a peace of mind if they aren’t carrier or allow them to fully consider the risks and other options available to them if they happen to be.

Figure 1: If two people who are both carriers of a recessive mutation have an offspring, there is a risk that their child could receive both mutated versions of the gene and so have the genetic disease. Figure created on BioRender.
One option for carrier parents is preimplantation testing (See Figure 2), whereby eggs are fertilized in vitro – meaning in a laboratory – and then tested to see if they contain two copies of the recessive mutated gene. Only the fertilised eggs that don’t contain the mutations are then implanted into the woman’s uterus.
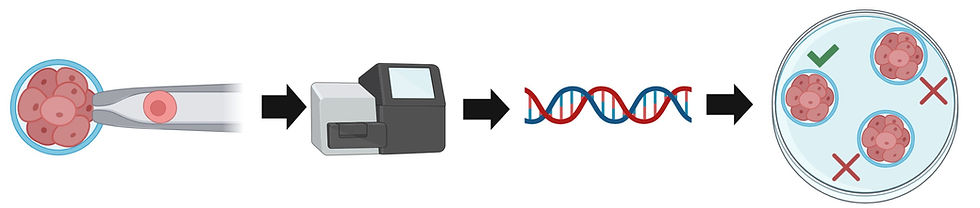
Figure 2: An overview of the preimplantation testing process. Eggs are extracted from a female and fertilised in vitro (i.e. in the lab) by sperm. They are then allowed to develop into embryos in vitro. After that, a cell from each embryo is extracted a genetically tested for a genetic disease their parents may be carriers of. At the moment, only the section(s) of DNA potentially containing the specific disease-causing mutation is sequenced. Sequencing is usually carrier out using a second-generation technique, such as Illumina. After sequencing each embryo, only one without a mutation will be implanted back into the mother’s uterus. Figure created on BioRender.
Why bother: the benefits of genetic testing
In addition to testing embryos, foetuses and newborns, adults at risk of or potentially already suffering from genetic diseases can be tested.
In comparison to the typical clinical diagnosis, clinical genetics may allow a more specific diagnosis, for example by distinguishing between diseases which have similar symptoms but different DNA mutations that cause them. Moving forward, if these diseases respond differently to treatment, this can also lead to more specific and effective treatment plans.
Genetic testing can be more cost effective in the long-term despite its high price tag by preventing unnecessary further testing and check-ups for healthy patients. It can speed up the diagnostic process, which can prevent the need for blindly conducting trials and errors of many different treatments, some of which could even be unknowingly harmful or counterproductive.
Finding mutations (a.k.a. the needle in a haystack)
The DNA variation causing many genetic conditions are still unknown. Finding them is made harder by the fact that genetic disorders can be caused by multiple types of DNA variation. For example, some are caused by as little as a single base mutation, such as Huntington disease, whilst others are caused by an increase in the number of copies of a DNA sequence, or even by large deletions of parts of gene. Worse still, many genetic conditions are complex traits. This means that the risk of getting them depends on both the environment and multiple genetic mutations. Interpreting data to find all the significant mutations is much harder for complex traits than for simple ones, which only depend on the sequence of one gene.
Another complication to finding disease-causing mutations is that some mutations show incomplete penetrance. This means that the mutation may cause the disease in some individuals and not in others. This makes interpreting its inheritance patterns and determining its significance very difficult.
If a specific mutation is found more often in people with a genetic disease than in healthy people, it may contribute to the disease. Deciding this involves checking previous research on the genes and proteins the mutation effects, as well as a lot of complex interpretation and statistics. For example, we can look into whether the mutation changes a proteins amino acid sequence and/or activity. This can be investigated in a laboratory or initially modelled on the computer, which can be faster and cheaper. The mutation must also show a pattern of inheritance expected for genetic diseases before it can eventually be classified.
The American College of Medical Genetics had a widely used classification system in which mutations are grouped into five classes: benign, likely benign, uncertain, likely pathogenic, and pathogenic. Classes 4 and 5 are able to be used for diagnostic testing.
However, as sequencing becomes cheaper, more and more information about disease-associated mutations is being discovered. Mutations in lower classes may be moved up into classes 4 or 5 based on new information. This shows the importance of research using larger groups of people and of re-evaluation of old information, especially when patients’ health is at stake.
How it is done - a brief introduction to sequencing
Original DNA sequencing technology was slow and expensive. Currently genes are usually sequenced using ‘second generation’ technology (Check out Table 1). This involves creating many copies of the DNA to be sequenced, and then cutting it into many short fragments to be sequenced.
Sequencing is usually automated, whereby the process begins with the DNA being unwound and split into single strands. Then, the complimentary strand is remade using different fluorescent labels for each base. This technology works best with only short strands of DNA, and it can become costly and laborious to ‘stich together’ many strands if you want to sequence a larger section of DNA.
Recently there has been an increase in whole genome sequencing, in which all a person’s DNA is sequenced and analysed. This is because of improvements in sequencing technology, meaning that it is cheaper and quicker than ever before to sequence DNA. This so called ‘third generation’ sequencing often only requires an extremely small sample and can sequence much longer DNA fragments. However, its accuracy is still low, and the technology still required further refinement. The amount of data stored in the whole genome is vast, and the significance of many mutations found is still unknown.
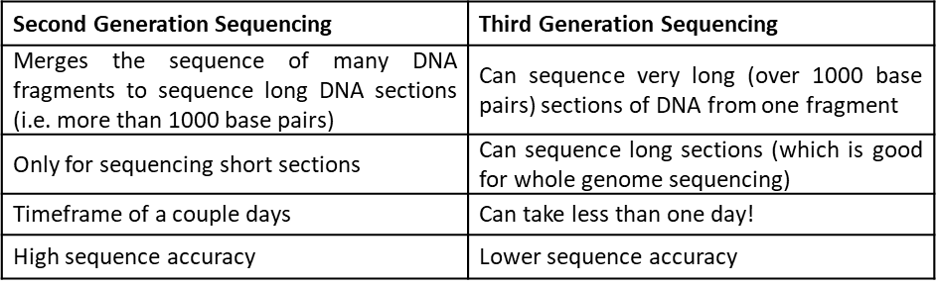
Table 1: A comparison between second- and third- generation sequencing.
Future trends: moving towards whole genome sequencing
At the moment, tests for a specific disease condition will only sequence the section of DNA containing the possible mutation. This means that these tests for some conditions will miss out on detecting other conditions. In addition, testing for many different conditions is time consuming and expensive.
Therefore, cheap and quick whole genome sequencing could be the answer to this problem as it could detect all types of genetic variation. However, as sequencing costs continue to decrease, the amount of sequences needed to be interpreted continues to rise. Interpreting and finding specific mutations in long DNA sequences can be a challenge in itself. As whole genome sequencing becomes more accessible, the problem of interpreting the result will only magnify.
The original Human Genome Project (HGP) , completed in April 2003, was the first full sequencing of a human’s genome. It was an international effort, costing around $2.7 billion and taking years to complete. Technology developed and improved during the HGP and onwards has allowed improvements in sequencing speed and accuracy. This trend continued with the more recent 100,000 genomes project, which aimed to sequence 100,000 genomes from the National Health Service (NHS) patients with rare diseases or cancer. The sequences from this project are still being used to research into the genetics of rare diseases, helping to find their causes and new treatments. This was completed in 2018. Now it can cost as little as around $1000 to sequence a genome in a matter of days!
If these trends continue, whole genome sequencing will become cheaper and cheaper. This makes whole genome sequencing easily scalable. Another benefit is that using the result of this one test, many different mutations causing different conditions can be searched for. Perhaps in the future we will all have our genomes sequenced and screened for diseases!
However, whole genome sequencing has its own risks. Such large amounts of data mean that if it is not handled correctly, small but crucial mutations could be lost or misinterpreted. Also, whole genome sequencing cannot detect DNA variation that isn’t related to sequence; many diseases are caused by variation in DNA structure.
From diagnosis to treatment: pharmacogenetics
A patient’s genetics can positively or negatively impact their response to certain drugs. Pharmacogenetics studies this so that patients can be grouped based on their response. This makes treatment safer, as a drug will only be used for patients who find the drug safe and effective. This can also save healthcare providers money as they aren’t wasting drugs on people who are unlikely to respond to it.
Pharmacogenetics is still a relatively new subject. In the UK, the National Health Service only recently began implementing pharmacogenetic testing to try and minimise the toxic side effects of cancer treatments for patients. This has the possibility to revolutionise the way we prescribe medicine to people. Initially, pharmocagenetics will likely focus on treatments with clear-cut responsed in patients and toxic side effects. In the future, as more information is gathered and accuracy improves, a quick genetic test before taking prescription drugs could become the norm!
The ethical controversy of clinical genetics
The ethics surrounding genetic testing can be controversial and requires explicit legislation. The full understanding and consent of the participant, including an understanding of the possible risks and consequences, must be received before the test is carried out, which is also known as informed consent. Genetic counsellors are, therefore, an important part of a clinical genetics team, as they help guide people through this decision. They also help patients understand their results.
The main ethical issue surrounding gene sequencing is who should own and/or benefit from the sequencing. Should companies be allowed to profit from sequencing patients who did not get paid themselves? Another main issue surrounds privacy – if you have had your genome sequences, who should have access to it? The data could be valuable to other researchers and help make new discoveries, but it then could also be used in ways you didn’t originally want it to be. Patients involved in the 100,000 genomes project had to give informed consent to be involved and were asked if they were okay with companies using the data from their genome for research, and the research projects undergo strict ethical reviews before approval. Furthermore, any data used is anonymous and cannot be traced back by researchers to specific people. Genomics England, which was set up by the UK government to run the 100,000 genomes project, has an independent Ethics Committee to oversee and advise on ethical concerns.
Many people worry that if genetic data was public, those with mutations associated with diseases may have increased insurance costs or be treated differently by an employer. This is called genetic discrimination. In the UK, employers would not be allowed by law to ask for genetic information due to the Equalities Act, but many say this is not enough, and for example it does not prevent requests from insurance providers. Laws in the United States, specifically the Genetic Information Non-discrimination Act (GINA), also prevent genetic discrimination.
Ultimately, improvements in genetic sequencing and other technologies now allows genetic testing on a larger scale than ever before, and with a greater level of specificity. These genetic tests can be invaluable in the clinic to help diagnose rare diseases and can help doctors decide which treatment plan is most likely to work.
There are more than 1000 genetic tests currently available, but choosing which to use, and how to interpret them, can be difficult. Clinical geneticists are working towards a future where there will be one low cost, easily accessible WGS test which can detect all types of disease-causing sequence mutations. This will revolutionise patient care but making sure patient privacy does not pay the price will take some careful planning.
Author: Rose Begg, BSc Biochemistry
Disclaimer: All figures created using BioRender are intended solely for educational purposes and not for profit.



